 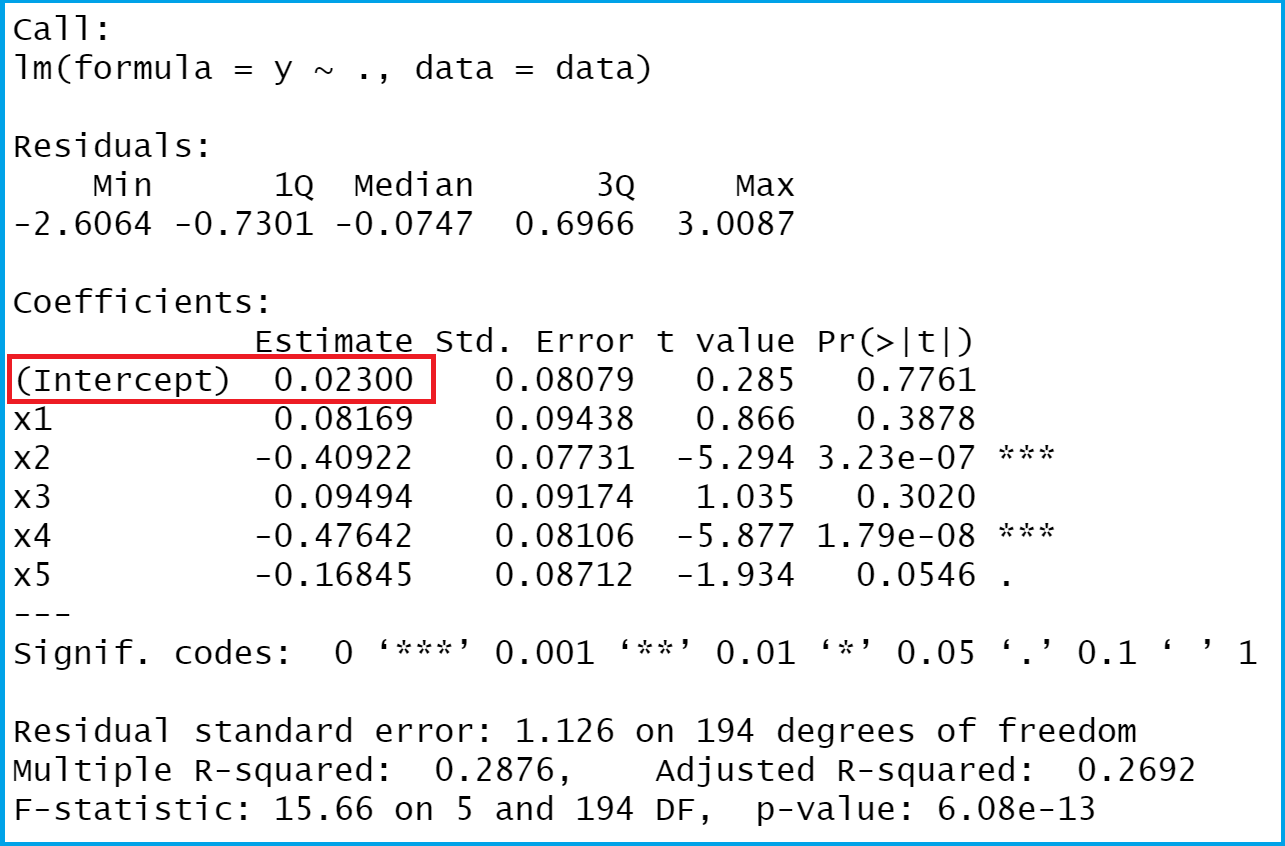
We will explore this issue later in this lab. There is some evidence for non-linearity in the relationship between lstat and medv. We will now plot medv and lstat along with the least squares regression line using the plot() and abline() functions. As expected, the confidence and prediction intervals are centered around the same point (a predicted value of 25.05 for medv when lstat equals 10), but the latter are substantially wider. The predict() function can be used to produce confidence intervals and prediction intervals for the prediction of medv for a given value of lstat. In order to obtain a confidence interval for the coefficient estimates, we can use the confint() command. "qr" "df.residual" "xlevels" "call" "terms" "model" coef(lm.fit) (Intercept) lstat names(lm.fit) "coefficients" "residuals" "effects" "rank" "fitted.values" "assign" lm.fit$coefficients-it is safer to use the extractor functions like coef() to access them. Although we can extract these quantities by name-e.g. We can use the names() function in order to find out what other pieces of information are stored in lm.fit. Residual standard error: 6.216 on 504 degrees of freedom This gives us pvalues and standard errors for the coefficients, as well as the \(R^2\) statistic and \(F\)-statistic for the model. For more detailed information, we use summary(lm.fit). If we type lm.fit, some basic information about the model is output. If we attach Boston, the first line works fine because R now recognizes the variables. The next line tells R that the variables are in Boston. The command causes an error because R does not know where to find the variables medv and lstat. lm.fit=lm(medv~lstat) Error in eval(predvars, data, env) : object 'medv' not found The basic syntax is lm(y~x,data), where y is the response, x is the predictor, and data is the data set in which these two variables are kept. We will start by using the lm() function to fit a simple linear regression model, with medv as the response and lstat as the predictor. To find out more about the data set, we can type ?Boston. We will seek to predict medv using 13 predictors such as rm (average number of rooms per house), age (average age of houses), and lstat (percent of households with low socioeconomic status). 
The MASS library contains the Boston data set, which records medv (median house value) for 506 neighborhoods around Boston. However, the library() function must be called each time you wish to use a given package. This installation only needs to be done the first time you use a package. Alternatively, this can be done at the R command line via install.packages("ISLR"). Simply select the package you wish to install and R will automatically download the package. After you select any mirror site, a list of available packages will appear. For example, on a Windows system, select the Install package option under the Packages tab. However, other packages, such as ISLR, must be downloaded the first time they are used. Some libraries, such as MASS, come with R and do not need to be separately installed on your computer. If you receive an error message when loading any of these libraries, it likely indicates that the corresponding library has not yet been installed on your system. We also load the ISLR package, which includes the data sets associated with this book. Here we load the MASS package, which is a very large collection of data sets and functions. Basic functions that perform least squares linear regression and other simple analyses come standard with the base distribution, but more exotic functions require additional libraries. The library() function is used to load libraries, or groups of functions and data sets that are not included in the base R distribution. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
